Numerical precision of euclidean_distances with float32
See original GitHub issueDescription
I noticed that sklearn.metrics.pairwise.pairwise_distances function agrees with np.linalg.norm when using np.float64 arrays, but disagrees when using np.float32 arrays. See the code snippet below.
Steps/Code to Reproduce
import numpy as np
import scipy
import sklearn.metrics.pairwise
# create 64-bit vectors a and b that are very similar to each other
a_64 = np.array([61.221637725830078125, 71.60662841796875, -65.7512664794921875], dtype=np.float64)
b_64 = np.array([61.221637725830078125, 71.60894012451171875, -65.72847747802734375], dtype=np.float64)
# create 32-bit versions of a and b
a_32 = a_64.astype(np.float32)
b_32 = b_64.astype(np.float32)
# compute the distance from a to b using numpy, for both 64-bit and 32-bit
dist_64_np = np.array([np.linalg.norm(a_64 - b_64)], dtype=np.float64)
dist_32_np = np.array([np.linalg.norm(a_32 - b_32)], dtype=np.float32)
# compute the distance from a to b using sklearn, for both 64-bit and 32-bit
dist_64_sklearn = sklearn.metrics.pairwise.pairwise_distances([a_64], [b_64])
dist_32_sklearn = sklearn.metrics.pairwise.pairwise_distances([a_32], [b_32])
# note that the 64-bit sklearn results agree exactly with numpy, but the 32-bit results disagree
np.set_printoptions(precision=200)
print(dist_64_np)
print(dist_32_np)
print(dist_64_sklearn)
print(dist_32_sklearn)
Expected Results
I expect that the results from sklearn.metrics.pairwise.pairwise_distances would agree with np.linalg.norm for both 64-bit and 32-bit. In other words, I expect the following output:
[ 0.0229059506440019884643266578905240749008953571319580078125]
[ 0.02290595136582851409912109375]
[[ 0.0229059506440019884643266578905240749008953571319580078125]]
[[ 0.02290595136582851409912109375]]
Actual Results
The code snippet above produces the following output for me:
[ 0.0229059506440019884643266578905240749008953571319580078125]
[ 0.02290595136582851409912109375]
[[ 0.0229059506440019884643266578905240749008953571319580078125]]
[[ 0.03125]]
Versions
Darwin-16.6.0-x86_64-i386-64bit
('Python', '2.7.11 | 64-bit | (default, Jun 11 2016, 03:41:56) \n[GCC 4.2.1 Compatible Apple LLVM 6.0 (clang-600.0.57)]')
('NumPy', '1.11.3')
('SciPy', '0.19.0')
('Scikit-Learn', '0.18.1')
Issue Analytics
- State:
- Created 6 years ago
- Comments:102 (93 by maintainers)
 Top Results From Across the Web
Top Results From Across the Web
Numerical issues for alternative way to compute (squared ...
I am able to compute the distance matrix faster by a factor of ~10 compared to scipy.pdist with this. However, I observe numerical...
Read more >Fast and Numerically Stable Pairwise Distance Algorithms
It's not just that the results are sometimes negative but they are also sometimes equal up to float32 precision even if the points...
Read more >Euclidean Distance (Spatial Analyst)—ArcMap | Documentation
ArcGIS geoprocessing tool that calculates, for each cell, the Euclidean distance to the closest source.
Read more >Distance - Math.NET Numerics Documentation
float Hamming(Single[] a, Single[] b). Hamming Distance, i.e. the number of positions that have different values in the vectors.
Read more >sklearn.metrics.pairwise.euclidean_distances
Compute the distance matrix between each pair from a vector array X and Y. For efficiency reasons, the euclidean distance between a pair...
Read more > Top Related Medium Post
Top Related Medium Post
No results found
 Top Related StackOverflow Question
Top Related StackOverflow Question
No results found
 Troubleshoot Live Code
Troubleshoot Live Code
Lightrun enables developers to add logs, metrics and snapshots to live code - no restarts or redeploys required.
Start Free Top Related Reddit Thread
Top Related Reddit Thread
No results found
 Top Related Hackernoon Post
Top Related Hackernoon Post
No results found
 Top Related Tweet
Top Related Tweet
No results found
 Top Related Dev.to Post
Top Related Dev.to Post
No results found
 Top Related Hashnode Post
Top Related Hashnode Post
No results found

The raw values from the real world rarely have that kind of accuracy, that’s right. But ML isn’t limited to that kind of input. One might want to apply ML to mathematical problems, like applying MDS on the graph of a rubik’s cube-like puzzle or clustering the successful strategies found by your swarm of RL agents playing pacman. Even if the initial source of the information is the real world, there might be some mid-way processing that makes most digits relevant to the clustering algorithm. Like the result of a gradient descent on a function whose parameters are statistically sampled in the real world.
I’m actually wondering why we’re still discussing this. I guess we all agree that scikit-learn should try its best in the trade-off accuracy vs. computation time. And whoever isn’t happy with the current state should submit a pull request.
Here are some benchmarks for speed comparison between scipy and sklearn. The benchmarks compare
sklearn.metrics.pairwise.euclidean_distances(X,X)withscipy.spatial.distance.cdist(X,X)for Xs of all sizes. Number of samples goes from 2⁴ (16) to 2¹³ (8192), and number of features goes from 2⁰ (1) to 2¹³ (8192).The value in each cell is the speedup of sklearn vs scipy, i.e. below 1 sklearn is slower and above 1 sklearn is faster.
The first benchmark is using the MKL implementation of BLAS and a single core.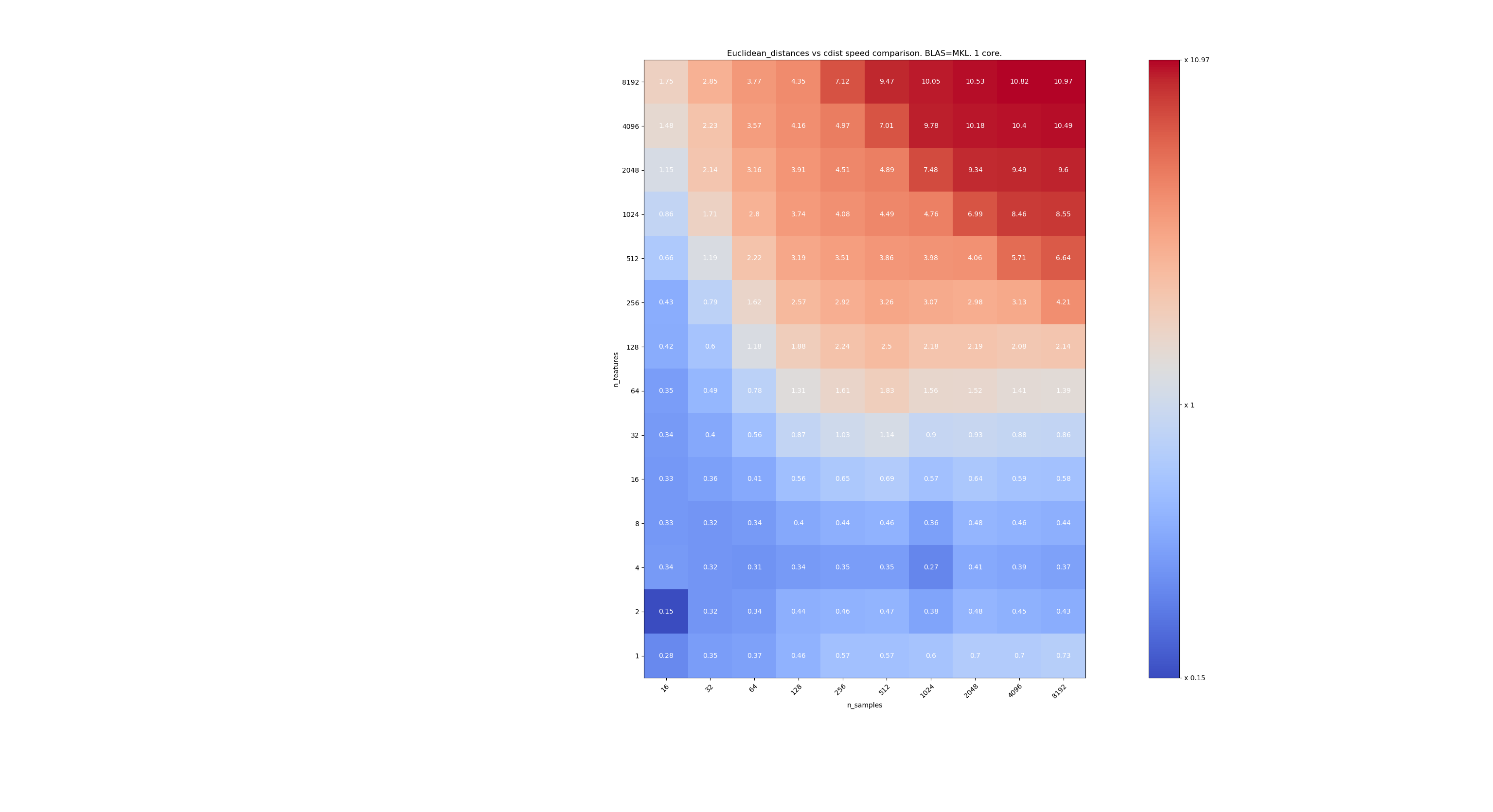
The second one is using the OpenBLAS implementation of BLAS and a single core. It’s just to check that both MKL and OpenBLAS have the same behavior.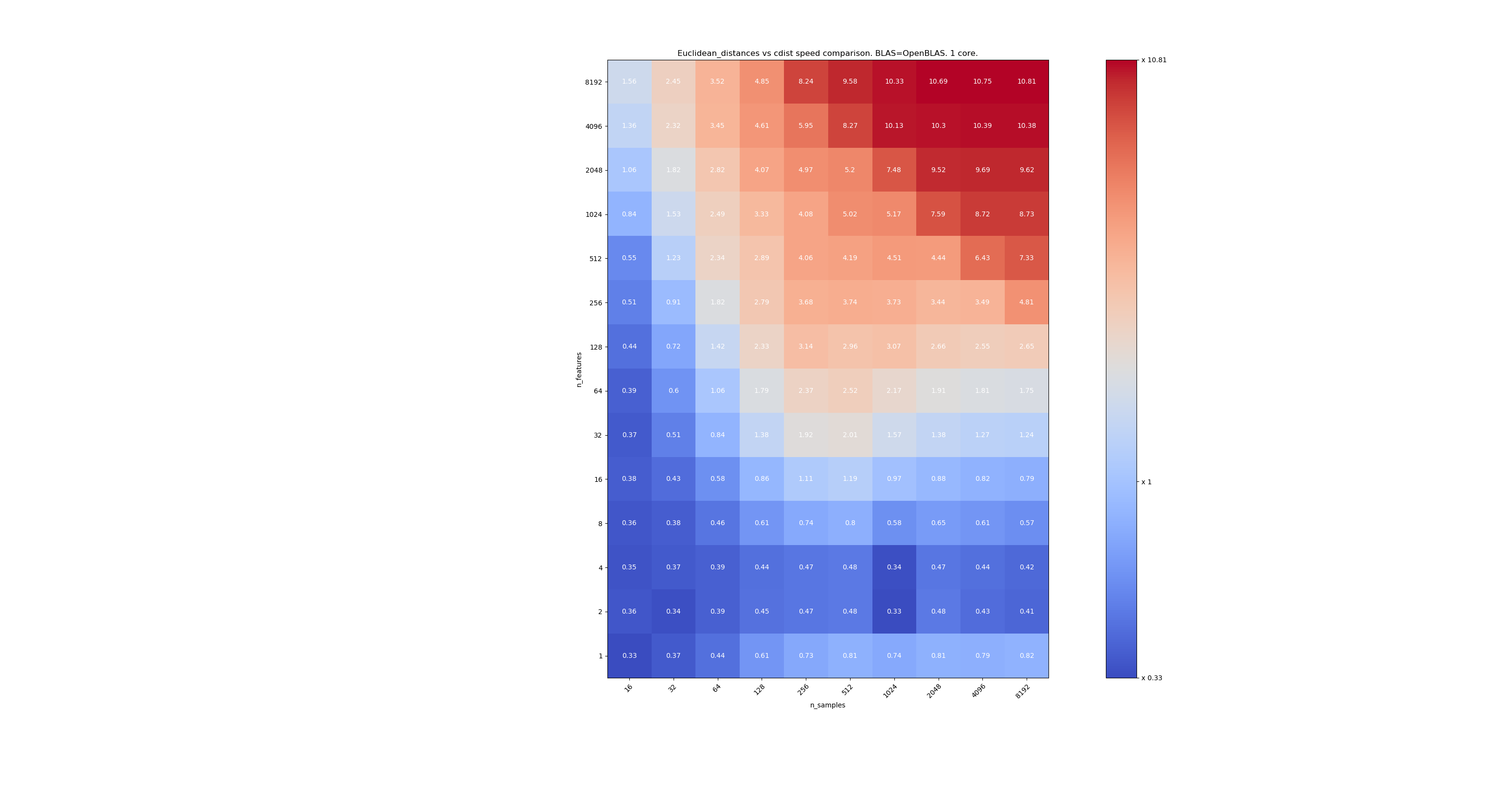
The third one is using the MKL implementation of BLAS and 4 cores. The thing is that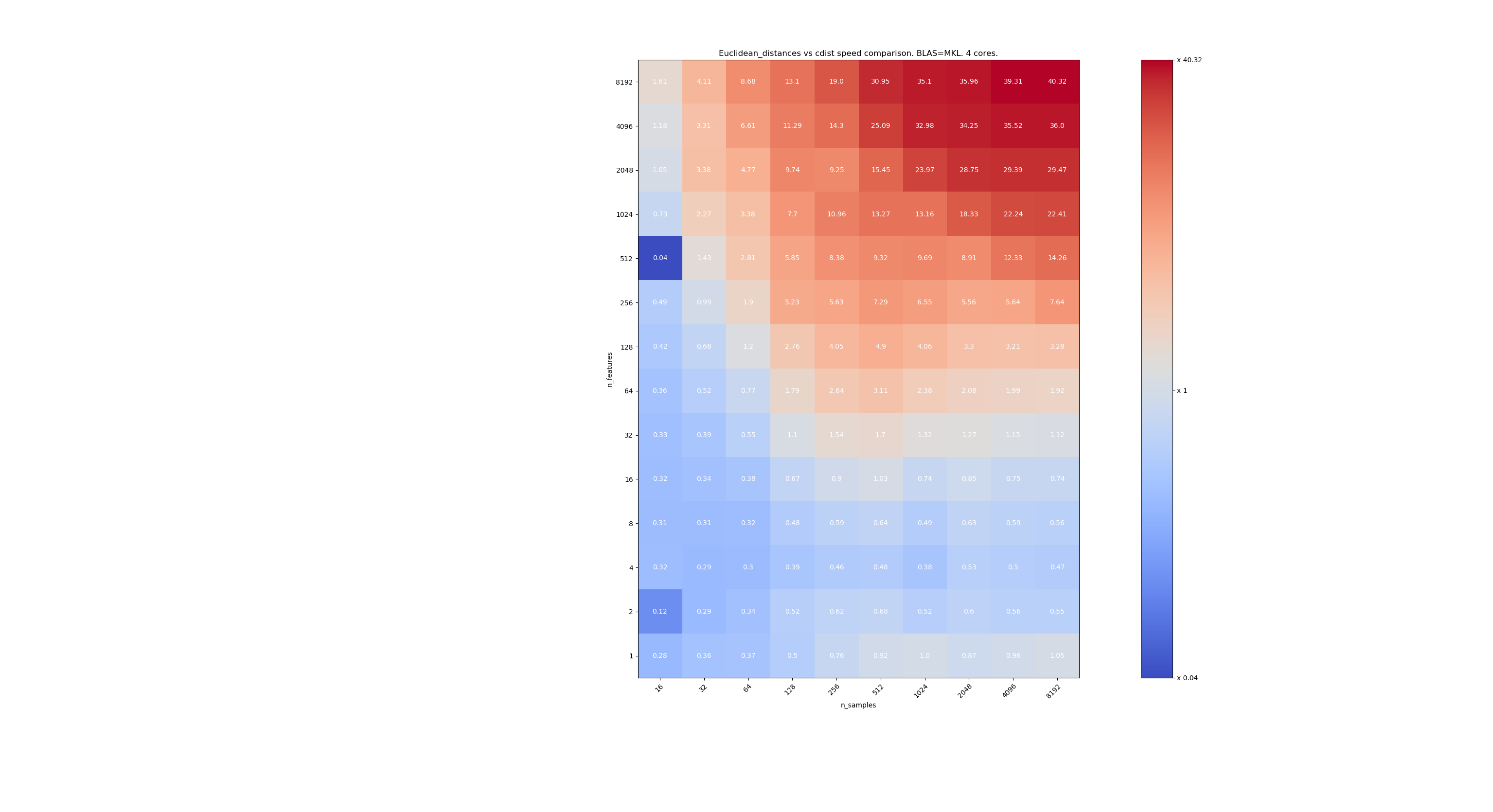
euclidean_distancesis parallelized through a BLAS LEVEL 3 function butcdistonly uses a BLAS LEVEL 1 function. Interestingly it almost doesn’t change the frontier.When n_samples is not too low (>100), it seems that the frontier is around 32 features. We could decide to use cdist when n_features < 32 and euclidean_distances when n_features > 32. This is faster and there no precision issue. This also has the advantage that when n_features is small, the julia threshold leads to a lot of re-computations. Using cdist avoids that.
When n_features > 32, we can keep the
euclidean_distancesimplementation, updated with the julia threshold. Adding the threshold shouldn’t sloweuclidean_distancestoo much because the number of features is high enough so that only a few re-computations are necessary.